AI tool helps match enzymes to substrates

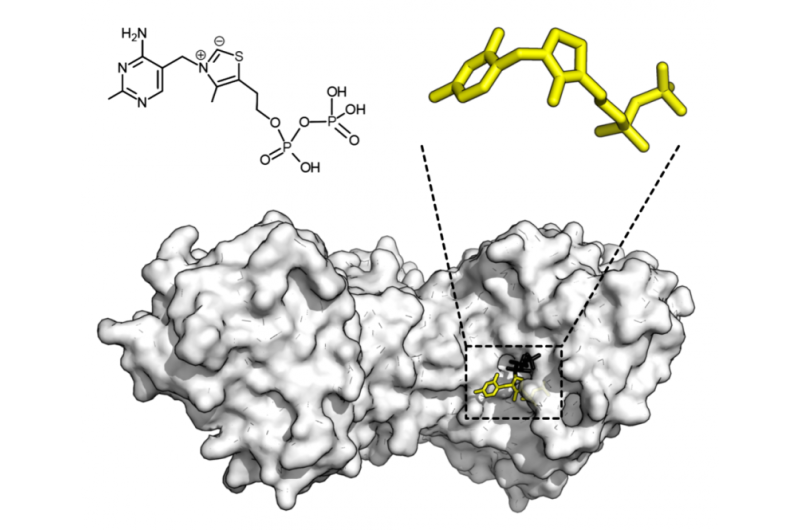
Chemical structure of thiamine pyrophosphate and protein structure of transketolase. Thiamine pyrophosphate cofactor in yellow and xylulose 5-phosphate substrate in black. Credit: Thomas Shafee/Wikipedia
A new artificial intelligence-based tool can help researchers determine how well an enzyme fits a desired target, helping them find the best combination of enzyme and substrate for applications ranging from catalysis to medicine to manufacturing.
Led by Huimin Zhao, professor of chemical and biomolecular engineering at the University of Illinois Urbana-Champaign, the researchers developed EZSpecificity using new data on enzyme-substrate pairs and a new machine learning algorithm. They made the tool freely available online and published their results in the journal Nature.
“If we want a certain product to use an enzyme, we want to use the best combination of enzyme and substrate,” said Zhao, who is also director of the NSF Molecule Maker Lab Institute and the NSF iBioFoundry at the U of I. “EZSpecificity is an AI model that can analyze an enzyme sequence and then predict which substrate can best fit that enzyme. It is highly complementary to the CLEAN AI model that we developed to predict the function of an enzyme from its sequence over two years ago.
Enzymes are large proteins that catalyze molecular reactions. They have pocket-like regions into which target molecules, called substrates, fit. The match between an enzyme and a substrate is called specificity. The typical analogy of enzyme-substrate interaction is that of a lock and key: only the right key will open the lock. However, the function of enzymes is not so simple, Zhao said.
“It’s difficult to find the best combination because the pocket is not static,” he said. “The enzyme actually changes its conformation when it interacts with the substrate. It’s more of an induced fit. And some enzymes are confusing and can catalyze different types of reactions. This makes prediction very difficult. That’s why we need a machine learning model and experimental data that actually proves which pairing will work best.”
Although other models of enzyme specificity have been introduced, their accuracy and the types of enzymatic reactions they can predict are limited.
Zhao’s group realized that to improve AI’s ability to predict specificity, they needed to improve and expand the dataset that the machine learning model draws inspiration from. They teamed up with the group led by Diwakar Shukla, a professor of chemical and biomolecular engineering at the U of I. Shukla’s group performed docking studies for different classes of enzymes to create a large database containing information not only about the sequence and structure of an enzyme, but also about how enzymes from different classes conform around different types of substrates.
“Experiments that capture how enzymes interact with their substrates are often slow and complex, so we performed extensive docking simulations to complement and expand existing experimental data,” Shukla said. “We zoomed in on the atomic-level interactions between enzymes and their substrates. Millions of docking calculations gave us this missing piece of the puzzle to build a highly accurate enzyme specificity predictor.”
The researchers then tested EZSpecificity side-by-side with ESP, the current leading model, in four scenarios designed to mimic real-world applications. EZSpecificity outperformed ESP in all scenarios. Finally, the researchers experimentally validated EZSpecificity by examining eight halogenase enzymes, a class that has not been well characterized but is increasingly used to make bioactive molecules, and 78 substrates. EZSpecificity achieved 91.7% accuracy for its best match predictions, while ESP only had 58.3% accuracy.
“I can’t say it works for all enzymes, but for some enzymes we showed that EZSpecificity actually works very well,” Zhao said. “We want to make this tool accessible to others, so we developed a user interface. Researchers can now input the substrate and protein sequence and then use our tool to predict whether that substrate may work well or not.”
Next, the researchers plan to expand their AI tools to analyze enzyme selectivity, which indicates whether an enzyme has a preference for a certain site on a substrate, to help rule out enzymes with off-target effects. They also plan to continue refining EZSpecificity with more experimental data.
More information:
Haiyang Cui et al, Prediction of enzyme specificity using cross-attention graph neural networks, Nature (2025). DOI: 10.1038/s41586-025-09697-2
Provided by University of Illinois at Urbana-Champaign
Quote: AI tool helps match enzymes to substrates (October 8, 2025) retrieved October 8, 2025 from https://phys.org/news/2025-10-ai-tool-enzymes-substrates.html
This document is subject to copyright. Except for fair use for private study or research purposes, no part may be reproduced without written permission. The content is provided for informational purposes only.

